DEWE (http://www.sing-group.org/dewe/) is an open source application for easily executing Differential Expression analyses in RNA-Seq data.
DEWE offers built-in, easy-to-configure workflows that facilitate the execution of DE analyses. Currently, DEWE provides two differential expression analysis workflows: HISAT2, StringTie and Ballgown and Bowtie2, StringTie and R libraries (Ballgown and edgeR).
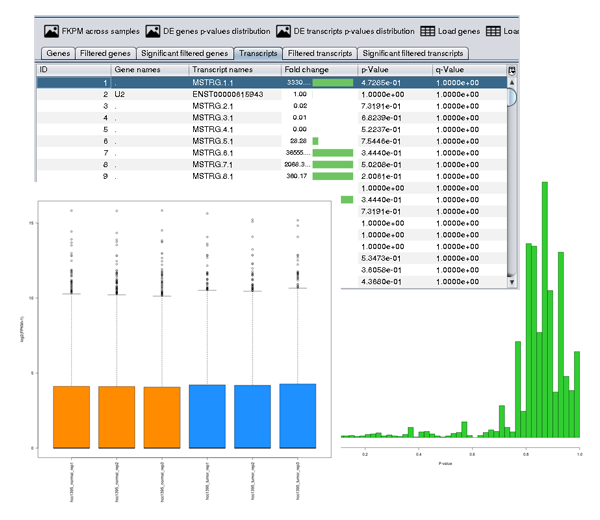
A key feature of DEWE is the interactive results management and visualisation. The main results of the execution of a built-in workflow are the outputs provided by Ballgown or edgeR. These outputs are automatically visualised in DEWE after the workflow execution and they can be reopened at any later time.
DEWE is available under two different installation methods: through a Docker container or through the use of a Virtual Box Machine. The Docker installers are available for the following operating systems: Windows 7 or higher, Linux with 3.10 kernel minimum and Mac OS X "Mountain Lion" or newer.