Similar to Evolutionary Trace, it should be possible to plot levels of selection (omega or a related number) on existing structures. But all I can find is SWAKK, which only allows two sequences, hence works with a window. I really just want to plot omega values of residues, calculated based on a >10 sequences MSA, on a structure. Does this exist? I am not into programming but it seems no too difficult, hence if it doesn´t exist I might ask my students to do this but if it already exists...Cheers
Have you looked at ConSurf server or ConSurfDB ?
Consurf uses Rate4Site to derive phylogenetic tree of the homologues using the neighbor joining algorithm (using the JTT substitution model).
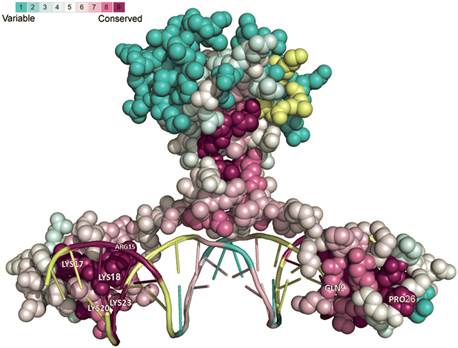
Hi Khader
Tx for you reply. Consurf is similar to Evolutionary Trace and shows variation at the amino acid level. Typically you need or at least use divergent datasets and you look at divergence (hence difference between species). What I want is for instance to determine which part of a protein is under positive selection. DNAsp can determine which stretches of sequences are under positive selection and yields basically a numerical output, this I want to plot on a PDB strcuture. For this type of analysis you use nucleotide sequences of strains or varieties, hence you are looking at polymorhisms (differences within spècies level) and you calculate the ratio of non-synonymous to synonymous mutations. Such that you have an idea: the sequences you use are typically 95-98% identical. Consurf cannot do anything with this. In reality right now I will be helped by simply plotting the data on a PDB, in the future I foresee I want to screen in 3D space but that is way more complicated!!
Use of this site constitutes acceptance of our User Agreement and Privacy Policy.


AFAIK, Consurf works well with both nucleic acid sequence and amino acid sequences. The metric is similar to the approach you are mentioning (See the detailed methodology http://consurf.tau.ac.il/overview.html) but not exactly the same.